 
Its p K a does not belong to the predefined The group identified as ‘base2’īelongs to an N-terminus of a noncanonical amino acid therefore, The uncertainty of the calculated pI, which is reported as mean value The variationīetween different p K a sets allows judging The calculated pI when the latter is poorly defined. The isoelectric interval denotes the span of pH within which the predicted  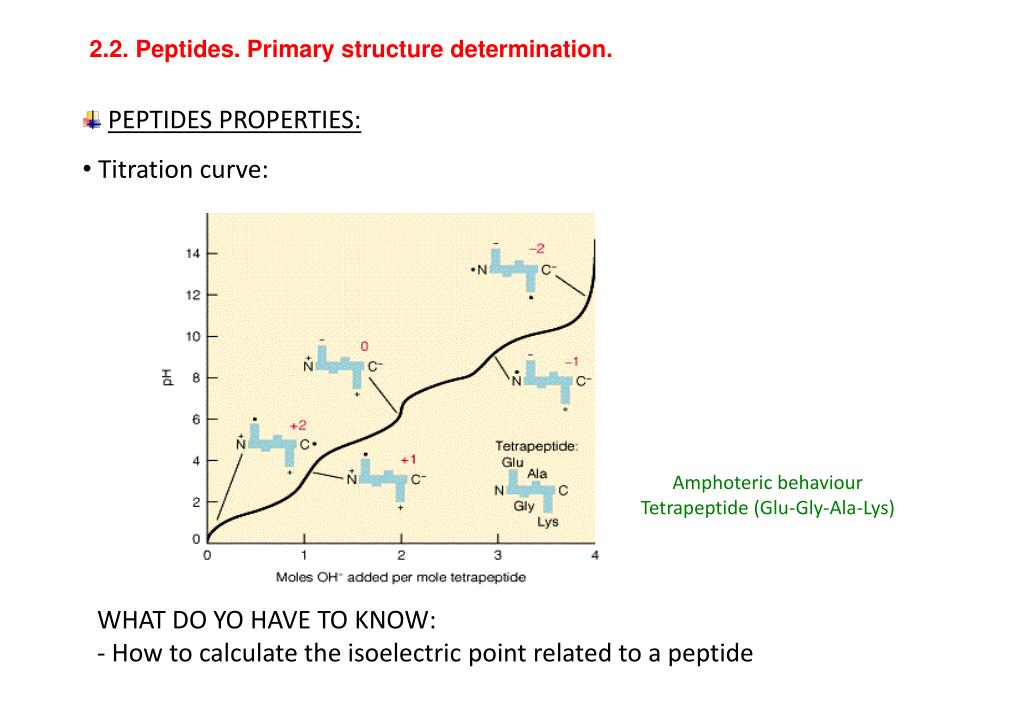
The white dot corresponds to the isoelectric point of the molecule. To the p K a values of the ionizable groupsįor the case of the IPC_peptide p K a set. P K a (base1), p K a (base2)) mark the inflection points, which correspond approximately To various predefined p K a sets of canonicalĪmino acids. (a) Charge versus pH curves resulting from the Henderson–HasselbalchĮquations for the dipeptide shown in (b). We anticipate that the pI calculation tool may facilitate optimization and purification activities across various application domains of peptides, including the development of biopharmaceuticals. The method is available in the form of an open source Python library at, which can be integrated into other proprietary and free software packages. The method is validated on a curated set of experimental measures on 29 modified and 119093 natural peptides, providing an improvement of R 2 from 0.74 to 0.95 and 0.96 against the conventional sequence-based approach for modified peptides for the two studied p K a prediction tools, ACDlabs and pKaMatcher, correspondingly. In this work, we describe a pI calculation method that uses the chemical structure as an input, automatically identifies ionizable groups of nCAAs and other fragments, and performs p K a predictions for them. Noncanonical modifications (nCAAs) are ignored in the conventional sequence-based pI calculations, therefore producing large errors in their pI predictions. 
Different p K a sets are published in the literature for natural amino acids, however, they are insufficient to describe synthetically modified peptides, complex peptides of natural origin, and peptides conjugated with structures of other modalities. Experimental measurements of pI can be replaced by calculations knowing the ionizable groups of peptides and their corresponding p K a values. It is widely used to steer design away from low solubility and aggregation and guide peptide separation and purification. The isoelectric point (pI) is a fundamental physicochemical property of peptides and proteins. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
